Oxygen Sensing
OXYGEN SENSING – how the kidney and nephrologists contributed to illuminating a fundamental biological process
by Chris Pugh
Executive summary
In the text below I narrate a personal view of work in Oxford led by Peter Ratcliffe that led to his receipt of a share in the 2019 Nobel Prize in Physiology or Medicine “for discoveries of how cells sense and adapt to oxygen availability”. The article outlines the journey from identification of an oxygen regulated enhancer element 3’ to the mouse erythropoietin gene, through the important recognition that this enhancer operated, and regulated other genes, in multiple cell types (that do not make erythropoietin) across metazoan species. The cognate transcription factor, HIF, was identified by Gregg Semenza at Johns Hopkins, but the Oxford group defined how its abundance and activity was controlled by a family of oxygen-consuming ‘HIF-hydroxylase’ enzymes. This work also defined the function of the protein mutated in von Hippel Lindau disease and almost all spontaneously occurring renal clear cell cancers. In parallel the group in Oxford also unequivocally defined the erythropoietin producing cells in kidney and liver.
The HIF transcription factors and the controlling ‘HIF-hydroxylase’ enzymes have been targeted by drugs now licensed for clinical use. At present, the HIF-hydroxylase inhibitors (e.g. Daprodustat, Roxadustat, Vadadustat) are available to treat renal anaemia and Belzutifan, that targets HIF-2 dimerisation, is used in cancer treatment. However, given the wide action of this system it seems likely that carefully timed and targeted administration of drugs that target this pathway will have wider applications, for example in ischaemic and metabolic conditions.
The Story
Anyone who has been trekking in the Himalayas knows the importance of acclimatisation, the process by which the body adapts to maintain its ability to function despite the reduced oxygen levels present at altitude. One of the most striking features of this acclimatisation is an increase in the concentration of erythrocytes in the blood, thereby increasing the amount of oxygen that can be carried per litre, but many other adaptations occur in parallel, including effects on breathing, the circulation and metabolism. How does this occur in a co-ordinated way? What is its medical relevance? What does it have to do with the kidney? Read on and hopefully the answers will become apparent!
In 1982 I had just completed a doctorate in Immunology and entered the clinical school at Oxford. I was one of a group of about twelve medical students attached to the Nuffield Department of Medicine where the general medical registrar was Dr. Peter Ratcliffe. I was initially attached to the Haematology side of the firm but I had seen enough of Peter to recognise him one weekend in the autumnal mud of the Oxford Sailing Club dinghy park looking slightly forlornly at a battleship grey Enterprise dinghy. We fell into conversation and it transpired that Fiona, then his fiancée, was on-call in London and he had nobody to sail with. I offered to crew for him and fortunately my sailing curriculum vitae was deemed adequate. Sailing together became a regular activity. Little did I know where this chance meeting and shared interest would go…..
Whilst I was starting to learn clinical medicine Peter had moved from the wards to the laboratory, working towards his Cambridge MD thesis on ‘Ischaemic and non-ischaemic factors in acute renal failure’. Isolated perfused rat kidneys were used to examine the nephrotoxicity of snake venoms (very damaging) and myoglobin (innocuous on its own). NMR spectroscopy was performed to assess the effects of haemorrhage on renal energetics in the anaesthetised rat. He also investigated the relationship between overall oxygen delivery and hypoxic cellular injury in isolated perfused rat kidneys. These experimental protocols were immensely challenging and associated frustrations were discussed in the car on the way to sailing or in the boat when the wind was light!
Somewhere on this journey I became interested in Nephrology as a specialty. As a medical student I did a brief attachment at the Oxford Renal Unit and was impressed by the range of clinical problems encountered. Des Oliver’s ability to combine application of technology and humanity to care coupled with an All Black team spirit and all conquering approach was impressive. As a house physician in the Nuffield Department of Medicine I had the benefit of working on a firm in which John Ledingham was the consultant, Tony Raine the senior registrar and John Firth was the registrar. Whilst a renal SHO at the Hammersmith I benefitted from the support of Chris Winearls (amongst others) and had the privilege of presenting the experiences of Mrs IM, the first patient to experience a year of erythropoietin (Epo) therapy at the Staff Rounds, chaired inimitably by Keith Peters. Chris started a consultant post in Oxford in August 1988. I arrived as the new renal registrar in November; the senior registrar was none other than Peter Ratcliffe who was busy writing an application to the Wellcome Trust to study regulation of the Epo gene by hypoxia!

Peter Ratcliffe c1988
The grant application was successful and in 1989 Peter started two distinct projects, one attempting to use in vivo experiments and the isolated perfused kidney to understand the regulation of Epo gene expression and the other to attempt to identify, immortalise and grow Epo producing cells by generating transgenic mice that expressed SV40 large T antigen oncogene specifically in these cells. The rat work was undertaken in a laboratory on level 5 of the John Radcliffe Hospital whilst the molecular work took place on a bench in John Bell’s laboratory in the new Institute of Molecular Medicine (IMM). Progress on the first project was initially very frustrating. The then ‘in vogue’ technique for measuring RNA levels was Northern blotting which is of limited sensitivity and requires high quality preparations of intact full length RNA from tissues. Negative result followed negative result and the conclusion drawn was that the RNA was being degraded by ubiquitously present RNases. The quest led to increasingly lengthy protocols with ever more phenol/chloroform extraction steps; despite wearing double gloves and increasing treatments of glassware with DEPC nothing worked. Just as despair and superstitious ideas such as only working on moonless nights were beginning to creep in Peter was introduced to Richard Jones. Richard was a biochemist working on globin gene regulation who used an ‘RNase protection assay’ to measure RNA levels – this only depends on short fragments of RNA surviving extraction and it was possible to obtain meaningful results with brief protocols in which Richard indicated that gloves were only worn to protect against phenol burns and a cigarette could be smoked whilst doing the experiment (or at least in the gaps when centrifuges were spinning or gels were running)! A riboprobe for rat Epo mRNA was designed and results began to flow [Oxygen-dependent modulation of erythropoietin mRNA levels in isolated rat kidneys studied by RNase protection. Ratcliffe PJ, Jones RW, Phillips RE, Nicholls LG, Bell JI. J Exp Med. 1990 Aug 1;172(2):657-60. doi: 10.1084/jem.172.2.657.].
The second project was comparably frustrating. The idea was simple (and had precedence in the study of insulin producing pancreatic beta cells); use a large sequence of DNA from around the Epo locus to drive the expression of an oncogene in the cells that would normally make Epo, the oncogene would transform these cells and they would be easy to grow. Peter made a large DNA construct containing the SV40 large T antigen sequence inserted into the 5′ untranslated region of the mouse erythropoietin gene, flanked on each side by 9 and 7.5 kb of DNA from the mouse erythropoietin locus. He gave the construct to Martin Johnson in the Cambridge Anatomy Department to perform pronuclear injection. The creation of transgenic mice took time….. These mice ultimately proved valuable in a variety of ways which we will return to, but the original goal for which they were created, to immortalise and grow Epo producing cells, has still to be realised.
At this stage Peter was joined in the lab by a young Singaporean renal physician, Tan Chorh Chuan (aka Steven) who had come to Oxford to undertake a doctorate. With the technical help of Kay Yeates, June Shultz and Lynn Nicholls, they had a very productive time studying Epo gene expression in vivo and in the isolated perfused kidney. This work evaluated the organ distribution of Epo mRNA in uraemic and control rats, showing its presence not only in kidney and liver, but also at lower levels in brain, testis, spleen and lung. They showed changes in the relative organ distribution of Epo mRNA as animals developed and aged and inducibility of Epo mRNA levels in multiple organs by relatively brief exposure to hypoxia. Pharmacological experiments indicated that oxygen-regulated Epo mRNA accumulation was not affected by inhibition of oxidative phosphorylation or through activation of cyclic AMP. This work benefitted from a strong collaborative link with Kai-Uwe Eckardt, Armin Kurtz and Christian Bauer at the Physiology Institute in Zurich, and this was strengthened further when Kai spent a year in Oxford.
In 1990 I joined the efforts, revived from clinical medicine by 3 weeks of competitive sailing in Hong Kong. Peter had now moved out of John Bell’s lab and into Room 317 of the IMM, a laboratory shared with Richard Jones, his post-doc Paul Campbell and a visiting French scientist, Carole Beaumont, although the isolated perfused kidney experiments were still being performed in a laboratory on Level 5 of the John Radcliffe Hospital. Together with Chorh Chuan, Lynn, Kay and June we formed quite an eclectic interdisciplinary mix.

Peter Ratcliffe, Tan Chorh Chuan and Chris Pugh, Lubeck Erythropoietin meeting 1991.
I was supported by an MRC Training Fellowship even though I already held a doctorate. Reviewing the application now I note no lack of ambition – it states ‘The proposed project is intended to elucidate the mechanism by which changes in oxygen delivery are sensed by the kidney at the whole organ, cellular and ultimately molecular levels.’ Although I was originally going to join attempts being made to investigate mechanisms underlying the oxygen sensitivity of Epo mRNA production in the isolated perfused kidney it was clear that this system had many limitations for any form of precise dissection. Mark Goldberg and Frank Bunn at Harvard had recently shown that two human hepatoma cell lines, HepG2 and Hep3B, demonstrated oxygen-regulated Epo mRNA expression when the atmosphere above the tissue culture medium was severely oxygen-depleted. This indicated that all the necessary sensing and transduction components were present in these cells, making identification of all these necessary components potentially tractable and we set out to do this.
Firstly we sought to identify a DNA element close to the Epo gene that was necessary and sufficient for its transcriptional regulation by changes in oxygen levels. The approach was very simple. Peter and I used restriction endonucleases (enzymes that cut DNA at specific sequences) to prepare a variety of fragments from the mouse erythropoietin gene. We then inserted these pieces of DNA into one of the human hepatoma cell lines by ‘electroporation’. We used a home-made device that allowed a capacitor to be charged and then discharged through platinum electrodes inserted into the culture medium that contained both the cells and the pieces of DNA – it would probably get nowhere near passing current health and safety requirements! After the electric shock a high percentage of cells were dead but of those that survived a modest percentage would have taken in the DNA. The surviving cells were split into two culture flasks, one of which was then incubated overnight in normal oxygen levels (normoxia) and the other one placed in an incubator with low oxygen levels (hypoxia). The RNase protection assay was then used to measure the levels of mouse erythropoietin mRNA – if the DNA fragment contained the regulatory element there would be more Epo mRNA in the cells from the hypoxic flask than the normoxic flask. Many such regulatory elements are at the 5’ end of genes but as we nibbled away from that end all the fragments continued to show hypoxic upregulation. To investigate sections within, and downstream of, the gene we attached the fragments to an alpha globin gene and instead of looking for hypoxic regulation of Epo mRNA instead looked for regulation of the alpha globin reporter gene. Eventually I isolated a 123 bp Apa1-Pvu2 fragment situated at the 3’ end of the gene, just beyond the polyadenylation signal, that was sufficient to confer hypoxic regulation and had the characteristics of a ‘transcriptional enhancer’. Importantly, this element also conferred a response to cobaltous ions at concentrations known to stimulate Epo production, suggesting that both hypoxia and cobaltous ions worked in the same way to upregulate the Epo gene. Parallel work in laboratories in the USA identified a similar region in the human Epo gene and those studies were published during the period our own manuscript was being reviewed and ultimately published (June-December 1991) [Functional analysis of an oxygen-regulated transcriptional enhancer lying 3’ to the mouse erythropoietin gene. CW Pugh, CC Tan, RW Jones, PJ Ratcliffe. PNAS 88 (1991) 10553-10557. PMID: 1961720 PMCID: 52967].
IMAGE FOLLOWS SOON
Summary slide localising a hypoxically-regulated transcriptional enhancer in the mouse erythropoietin gene.
Upper panel shows a schematic of the mouse erythropoietin gene used in these experiments; the 5’ end is to the left, exons are shaded in grey and the Apa1-Pvu2 restriction fragment shaded in black.
Lower panel shows results of a RNase protection assay measuring relative mouse erythropoietin mRNA levels from cells incubated in normoxia (N) or hypoxia (H) following transfection with the entire DNA sequence (Intact), the DNA sequence with the Apa1-Pvu2 restriction fragment excised and the ends religated (Deleted) and this ‘deleted’ construct with the Apa1-Pvu2 restriction fragment reattached at the 5’ end (Repositioned); hypoxic induction of Epo mRNA is clearly dependent on the Apa1-Pvu2 restriction fragment, regardless of whether it is in its normal position or the alternative position.
At the end of this PNAS paper we tested the function of the DNA enhancer elements in two cell lines (CHO and MEL cells) that did not produce endogenous Epo and found no response to hypoxia in keeping with the then prevalent prejudice that the response of the erythropoietin gene to hypoxia was rather specific to this gene. In our rush to do these experiments we cultured the cells rather sparsely and did not take into account that the MEL cells grow in suspension rather than as a monolayer. I was concerned that this work was rather preliminary and might be a false negative result but despite these reservations the data was reported nonetheless.
Whilst this work was being undertaken Patrick Maxwell approached Peter, I think on Chris Winearls’ recommendation, about the possibility of doing a doctorate in the lab. We invited him to meet us one evening in my house in Headington. Patrick arrived late for what was effectively an interview dressed in a memorable turquoise track suit. He excused his lateness on the basis that since his last visit to the city someone had rather inconsiderately extended the M40 beyond Oxford without telling him and as he was rushing after a night on-call he had not noticed the Oxford turn and had carried on northwards towards Birmingham. Thankfully we were not put off by this slightly inauspicious start, which might well have debarred him from further consideration under current NHSE interview processes…. We didn’t enquire about his choice of clothing either but instead discussed the proposed work and his interest in it. Peter agreed to support him in preparing an application to the MRC for a Training Fellowship to work on the identification of the Epo producing cells in the kidney using the transgenic mice that expressed SV40 large T antigen under the control of the Epo locus.
Patrick’s application was successful and he joined us in the autumn of 1991. Transgenic mice bearing the Epo – SV40 large T antigen construct were eventually created in Cambridge and needed to be brought back to Oxford for analysis. Peter decided to fetch them in his car, quashing concerns about whether any welfare issues might arise by pointing out that the BMW he had acquired from his father was both comfortably air-conditioned and reliable! Once the mice were in Oxford a breeding colony needed to be established to generate a steady supply of animals for experimentation – this took time and meant Patrick was potentially underemployed. However, lots of other things were happening in parallel in a very interactive, collaborative and fun way. Lab notebooks from that period not infrequently have three, or four, people’s handwriting contributing different parts to the performance of a single experiment and Patrick joined these efforts.
We decided to revisit the question of whether the Epo hypoxic enhancer only worked in Epo-producing tissue culture cells. The culture conditions were adjusted and we gradually saw hints of hypoxic regulation in cells that did not express / regulate their endogenous Epo gene. Patrick methodically varied the cell density and growth conditions prior to transfection and saw increasingly robust responses from an eclectic mixture of cell types, including macrophages, skin and lung fibroblasts. As the work continued he increased the repertoire of cells showing a response and ultimately concluded it was ubiquitous, indicating that an oxygen-sensing system similar, or identical, to that controlling erythropoietin expression is wide-spread in mammalian cells. An important implication of this system persisting in cells that did not make erythropoietin was that it would regulate other genes in these cells in response to changes in oxygenation, although it was only later (and initially rather serendipitously, that we would identify these genes). Sadly, after a very protracted review process, the editors of Nature did not share our conviction that there was a widespread response which was of fundamental biological significance.

The work was eventually published some 18 months after the original submission to Nature in the Proceedings of the National Academy of Sciences of the USA [Inducible operation of the erythropoietin 3’ enhancer in multiple cell lines: evidence for a widespread oxygen-sensing mechanism. PH Maxwell, CW Pugh, PJ Ratcliffe. PNAS 90 (1993) 2423-2427. PMID: 8460154 PMCID: 46099].
IMAGE FOLLOWS SOON
Demonstration of hypoxically-regulated mouse erythropoietin 3’ transcriptional enhancer activity in multiple cell lines.
Results of a RNase protection assay measuring relative levels of FGH and alpha globin (a) from the four indicated cell lines incubated in normoxia (N) or hypoxia (H) following co-transfection with FGH (a plasmid constitutively expressing human growth hormone under the control of a mouse ferritin promoter) and either a plasmid containing an alpha globin gene coupled to the erythropoietin 3’ transcriptional enhancer (+) or not (-). In each case the cells transfected with the alpha globin gene coupled to the erythropoietin 3’ transcriptional enhancer (+) show a higher level of alpha globin mRNA when incubated in hypoxia than normoxia whereas this is not the case of the alpha globin gene on its own (-). This indicates that the enhancer is functioning not only in the Hep3B cells in which the endogenous Epo gene is known to be hypoxically regulated but also in U937, (a macrophage line) and MRC5 and IBR3 (lung and skin fibroblast cell lines) in which the endogenous Epo gene is not hypoxically regulated.
Having found an enhancer the next questions were obvious – what proteins bound to it and how was that binding controlled? As none of us were biochemists we decided against a traditional affinity purification approach (using the DNA sequence to fish out the proteins that bound to it from the mixture of proteins present in the nucleus of a hypoxic cell) and instead decided to try two approaches that fell under the general heading of ‘expression cloning’. As a step in this direction we refined the minimal DNA sequence necessary for hypoxic regulation and concatemerised it to maximise signal to noise. The general approach was then to connect this sequence to a detectable ‘reporter’ gene in a cell type that did not have the full regulatory system and then add a library of mammalian genes (fused to a transcriptional activator) to these cells, select those in which the reporter had become operative and retrieve the library component from these cells to identify the component that had restored function.
One expression cloning approach was undertaken using yeast, under the patient guidance of Rhona Borts and Ed Louis in the IMM. Yeast cells do make a transcriptional response to hypoxia but it does not utilise the enhancer sequence identified in the hepatoma cell lines, so in some ways they were ideal for identifying factors that might bind to this DNA sequence. Much work ultimately yielded two independent clones, 27.2 and 33.1 that bound to the mammalian DNA sequence but not to mutant sequences in a way that strongly mimicked the function of this sequence in mammalian cells. These clones were sequenced and found to both be derived from the same gene. Disappointingly, we were unable to make any sense of the function of this gene in mammalian cells, and still do not know what it does or why it can bind this DNA in yeast……
The second expression cloning approach was to use mammalian cells and was inspired by the work of Ian Kerr and George Stark at ICRF Lincoln’s Inn Fields and Dai Simmonds in Oxford, with whom we shared a larger laboratory on the top floor of the IMM. The original idea was to stably transfect cells with an artificial reporter transgene in which the enhancer sequence was linked to the gene for a cell surface antigen not normally present on those cells. Activation of the enhancer by hypoxia, or in normoxia by an incoming gene from a library based on transcripts made by mammalian cells exposed to hypoxia, would lead to expression of the cell surface antigen. These positive cells could then be retrieved using an antibody against that antigen, allowing recovery of the incoming gene responsible for enhancer activation. [A further refinement of this technique, which was to occupy Morwenna Wood and then Emma Vaux some years later for many, many hours was to mutagenize cells bearing this reporter construct, identify those that no longer responded to hypoxia, and then add libraries to complement the mutation, thereby allowing the gene to be identified]. The first step in this process was to make the reporter cells. HepG2 cells and Cos cells were each transfected with a mixture of a relatively large amount of DNA encoding our carefully designed reporter gene and a small amount of a plasmid encoding resistance to a toxic chemical. Doug Higgs kindly donated to us PGK-NeoR, a plasmid that he assured us would be widely expressed in cells and confer resistance to neomycin (or its analogue G418). On exposure to the toxin cells that had taken up PGK-NeoR would survive and because the reporter plasmid had been present in molar excess would almost certainly have integrated it too. Whilst this technique would only select cells in which the PGK-NeoR had integrated into open chromatin and would be expressed there was no guarantee that the reporter plasmid would also be in open chromatin and thus function. Frustratingly many clones were derived but at this stage none showed good reporter function. In trying to sort out what was going on and whether we had made a mistake in reporter gene design I discovered, much to my surprise, that although the reporter plasmid was not showing hypoxic regulation, the NeoR cassette was. Was the reporter gene enhancer working in trans or was PGK-NeoR itself hypoxia responsive? Rather shamefacedly I went to Doug Higgs to ask what PGK-NeoR stood for and he patiently explained that it was composed of a 506 base pair sequence from the phosphoglycerate kinase gene promoter fused to a neomycin resistance cassette. Peter and I immediately recognised that phosphoglycerate kinase was an enzyme in the glycolytic pathway and thus a prime candidate for a gene that might plausibly be upregulated in response to low oxygen levels. By chance we had stumbled on the first gene other than erythropoietin regulated by this system!
Around this time (1992-3) we had also been joined in the lab by John Firth, who planned to investigate endothelin gene expression, Ben Ebert, an American Rhodes Scholar who I had met when he was trying to identify a suitable project on which to undertake a doctorate and then Jonathan Gleadle, who also wished to do a DPhil. Together we made rapid progress using a hypothesis driven approach to identify other target genes regulated by this system, comparing the relevant DNA sequences and working on how they might be regulated [Oxygen-regulated control elements in the phosphoglycerate kinase 1 and lactate dehydrogenase A genes: similarities with the erythropoietin 3enhancer. JD Firth, BL Ebert, CW Pugh, PJ Ratcliffe. (1994) PNAS 91 6496-6500. PMID: 8022811 PMCID: 44229]. The number of genes regulated through this system was extended further by John O’Rourke, a post-doc employed on my MRC Clinician Scientist Fellowship, using an unprejudiced differential display PCR approach [O’Rourke JF, Pugh CW, Bartlett SM, Ratcliffe PJ. Identification of hypoxically inducible mRNAs in HeLa cells using differential-display PCR. Role of hypoxia-inducible factor-1. Eur J Biochem. 1996 Oct 15;241(2):403-10. doi: 10.1111/j.1432-1033.1996.00403.x. PMID: 8917436.]. More modern approaches now estimate the number of genes regulated by this system to be close to 1000.

A lab group (probably taken by Patrick Maxwell) in the Institute for Molecular Medicine, Oxford.
l-r Masaya Nagao, John O’Rourke, Sylvia Bartlett, Lynn Nicholls, Chris Pugh, Jonathan Gleadle, John Firth, Morwenna Wood, Kay Yeates, Peter Ratcliffe, Ben Ebert.

Peter Ratcliffe’s 40th birthday in my garden near The Kilns (home of CS Lewis) in Risinghurst (May 1994)
l-r Fiona Ratcliffe behind Patrick Maxwell, Peter Ratcliffe, Chris Dudley, Chris Winearls, John Firth


Different types of hard work and rest! (May 1995)
A parallel interest in the lab at the time was to identify whether an orthologous system existed in lower organisms – which would not only be of biological interest but which might provide more tractable systems in which to discover the key components – an idea which was to bear real fruit in 2001. We had drawn a blank in yeast, so moved to the fruit fly Drosophila melanogaster and subsequently the nematode worm C. elegans (of which more later). With Masaya Nagao, a visiting postdoc from Kyoto, Ben, Peter and I identified that drosophila cells contained a hypoxia-inducible complex that could bind to the mammalian erythropoietin enhancer element and that the drosophila phosphoglycerate kinase gene was hypoxically-regulated in these cells [Drosophila melanogaster SL2 cells contain a hypoxically inducible DNA binding complex which recognises mammalian HIF-binding sites M Nagao , B L Ebert, P J Ratcliffe, C W Pugh FEBS Letters 387 (1996) 161-6. PMID: 8674541]. This work was subsequently extended with Neil Bacon during his brief sojourn in the lab en route to founding Doctors.Net.UK! [Regulation of the Drosophila bHLH-PAS protein Sima by hypoxia: functional evidence for homology with mammalian HIF-1alpha. NCM Bacon, P Wappner, JF O’Rourke, SM Bartlett, B Shilo, CW Pugh, PJ Ratcliffe. Biochemical and Biophysical Research Communications 249 (1998) 811-816. PMID: 9731218].

Chris Pugh and Masaya Nagao (c1995)
Meanwhile, the mouse breeding programme had really picked up and Patrick took the lead on performing experiments to identify the specific cell types responsible for Epo production in the liver and kidney. At the time this was a controversial area with many believing the hormone was produced by tubular epithelial cells whilst others had indicated that Epo mRNA was largely in interstitial cells, although they were unclear whether these were fibroblasts or endothelial cells. Much to everyone’s surprise (Peter and Patrick’s total disbelief when I first mooted the hypothesis after looking at Patrick’s carefully performed Southern blots) the large DNA construct containing the SV40 large T antigen sequence inserted into the 5′ untranslated region of the mouse erythropoietin gene had homologously combined into the native Epo gene locus, meaning that (apart from any effects caused by the inserted DNA) the construct was subject to regulation by all the natural endogenous genetic influences, not just those encoded by the inserted transgene. Thus, there was no question that cells identified as expressing SV40 large T antigen by immunostaining would be the Epo producing cells. Peter’s isolated perfused kidney work had been heavily influenced by John Tange, a pathologist who was a perfectionist when it came to tissue fixation. This rubbed off on us and Patrick and I spent some initially frustrating, but ultimately satisfying, time devising the apparatus and acquiring the surgical skills to cannulate the infrarenal aorta of anaesthetised mice, adjust the pressure in the catheter to equate to the mouse’s mean arterial pressure and then cut the inferior vena cava allowing the kidneys to be perfused with fixative at this pressure (albeit without pulsatility). After processing, the fixed tissues were immunohistochemically stained for SV40 large T antigen and co-stained for markers of renal interstitial fibroblasts and other cell types. The resulting slides unequivocally identified the renal erythropoietin producing cells as interstitial fibroblasts [Identification of the renal erythropoietin-producing cells using transgenic mice. PH Maxwell, MK Osmond, CW Pugh, A Heryet, LG Nicholls, CC Tan, BG Doe, DJP Ferguson, MH Johnson, PJ Ratcliffe. Kidney International 44 (1993) 1149-1162. PMID: 8264149]. Parallel work revealed that in the liver Epo was produced by both hepatocytes and interstitial Ito cells [Expression of a homologously recombined erythropoietin-SV40T antigen fusion gene in mouse liver: evidence for erythropoietin production by Ito cells. PH Maxwell, DJP Ferguson, MK Osmond, CW Pugh, A Heryet, BG Doe, MH Johnson, PJ Ratcliffe. Blood 84 (1994) 1823-1830. PMID: 8080987], complementing the results of in situ hybridisation studies performed by others.
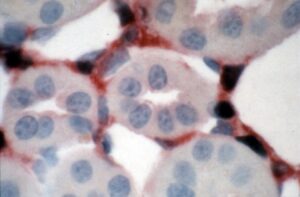 Picture of renal Epo producing cells (dark nuclei) co-labelled with an antibody to 5’ ectonucleotidase (red) a marker of renal interstitial fibroblasts.
Picture of renal Epo producing cells (dark nuclei) co-labelled with an antibody to 5’ ectonucleotidase (red) a marker of renal interstitial fibroblasts.

Patrick Maxwell and Mary Cotes in conversation during a walk at a conference (Ulm, Germany)
In 1993 the gene responsible for von Hippel-Lindau disease was cloned – I am not sure we noticed at the time…… More immediately significant for us, Gregg Semenza and Guang Wang at Johns Hopkins were making clear progress with the biochemical purification of the transcription factor that bound to the erythropoietin enhancer and in 1995 they described Hypoxia-inducible factor 1 as a basic-helix-loop-helix-PAS heterodimer in which a novel alpha chain bound to the already known about aryl hydrocarbon receptor nuclear translocator (ARNT) [Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension G L Wang , B H Jiang, E A Rue, G L Semenza Proc Natl Acad Sci U S A 1995 Jun 6;92(12):5510-4. doi: 10.1073/pnas.92.12.5510.].

Peter Ratcliffe and Gregg Semenza (Woods Hole, Massachusetts 1994).
We immediately wrote to Gregg, requesting the cDNA for HIF-1alpha and also contacted Oliver Hankinson, a British scientist who worked at UCLA on the aryl hydrocarbon receptor system, who very kindly provided us with a number of useful cell lines and reagents. With Oliver’s ARNT-deficient cells we were rapidly able to confirm the importance of this component for both hypoxic gene induction and tumour growth. Morwenna Wood, who had come to the lab after working as a renal SHO in Oxford, did much of this work whilst also setting up the mutagenesis and complementation project outlined earlier. [The role of the aryl hydrocarbon nuclear translocator (ARNT) in hypoxic induction of gene expression. Studies in ARNT deficient cells. SM Wood, JM Gleadle, CW Pugh, O Hankinson, PJ Ratcliffe. Journal of Biological Chemistry 271 (1996) 15117-15123. PMID: 8662957] [Hypoxia-inducible factor-1 modulates gene expression in solid tumors and influences both angiogenesis and tumor growth. PH Maxwell, GU Dachs, JM Gleadle, LG Nicholls, AL Harris, IJ Stratford, O Hankinson, CW Pugh, PJ Ratcliffe. PNAS 94 (1997) 8104-8109. PMID: 9223322 PMCID: 21564].
IMAGE FOLLOWS SOON
Comparison of tumours growing subcutaneously in mice following injection of ARNT competent (HIF+) and ARNT deficient (HIF-) mouse hepatoma cell lines
The right hand panel shows the more rapid growth of tumours derived from the ARNT competent cells. These larger tumours have an enhanced vascular network (third panel from left), induction of HIF-dependent genes (second panel from left shows in situ hybridisation for Glut-3 mRNA in the peri-necrotic (hypoxic) region) but despite these HIF-dependent adaptive changes the tumours contain larger areas of necrosis (N – in left hand panel) when compared with the tumours derived from ARNT deficient cells.

Chris Pugh, Patrick Maxwell and Morwenna Wood relaxing (and I am sure discussing science) c1996.

Chris Pugh and Morwenna Wood enjoying the products of some (culinary) biochemistry.
When we received the HIF-1alpha cDNA from Gregg we set about expressing protein fragments that we could use as immunogens to generate antibodies to allow protein detection. In parallel, we made DNA constructs to allow expression of fusion proteins that contained sections of HIF-1alpha linked to a heterologous DNA binding domain and in some cases a heterologous transactivation domain. These constructs allowed us to map different functional domains within the protein. We were able to show that the middle part of the protein contained a region that could confer oxygen-regulated changes in protein abundance on the fusion protein and the carboxy-terminal 50 amino acids of the protein contained a hypoxically-regulated transactivation domain [Activation of hypoxia-inducible factor-1; definition of regulatory domains within the alpha subunit. CW Pugh, JF O’Rourke, M Nagao, JM Gleadle, PJ Ratcliffe. Journal of Biological Chemistry 272 (1997) 11205-11214. PMID: 9111021]. Specifically, for HIF-1alpha we were able to show that when we made a fusion protein including amino acids 549-652 the level of protein was oxygen-regulated, whereas this was not true of a fusion protein including amino acids 572-652, implicating amino acids 549-571 as being necessary for this phenomenon. The prejudice at the time was that this type of post-translational regulation was generally mediated through amino acid phosphorylation but I mutated every single phosphoacceptor residue in this sequence without it having any significant impact on regulated protein stability. Out of shear curiosity I also mutated the two prolyl residues present and this did have a big effect – at the time we had no idea how this might be working so chose not to declare the result in our publication, fearing that it would raise more questions than it answered….
Around this time Peter, Patrick and I met Chris Schofield, following up on an interesting job advertisement I had spotted in Nature. Chris is an Oxford based chemist whose expertise would prove invaluable later in the story. However, at this stage although we had an interesting discussion about his work on the family of enzymes known as iron and oxoglutarate-dependent dioxygenases the link between our scientific interests was not apparent.
In further work we found a second region in HIF-1alpha able to confer similar effects and in fact both regions turned out to have highly similar primary amino acid sequences [Independent function of two destruction domains in hypoxia-inducible factor-alpha chains activated by prolyl hydroxylation. N Masson, C Willam, PH Maxwell, CW Pugh, PJ Ratcliffe. EMBO Journal 20 (2001) 5197-5206. PMID: 11566883 PMCID: 125617].
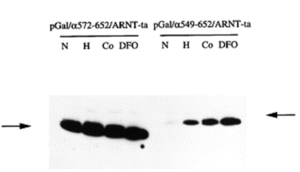
Amino acids 549-571 of human HIF-1 alpha are necessary for oxygen-regulated effects on protein abundance.Immunoblot illustrating that when expressed in cells the level of a fusion protein including amino acids 549-652 was oxygen-regulated (right panel), whereas this was not true of a fusion protein including amino acids 572-652 (left panel). Cells were exposed to normoxia (N), hypoxia (H) or medium containing cobaltous ions (Co) or desferrioxamine (DFO) – the latter three conditions being stimuli known to induce Epo expression in hepatoma cells).
Subsequently, a second HIF-alpha chain, originally named endothelial-PAS protein 1 (EPAS-1) was defined and we were able to compare its domain structure with that of HIF-1alpha [Oxygen-regulated and transactivating domains in endothelial PAS protein 1: comparison with hypoxia inducible factor-1alpha. JF O’Rourke, Y-M Tian, PJ Ratcliffe, CW Pugh. Journal of Biological Chemistry 274 (1999) 2060-2071. PMID: 9890965].
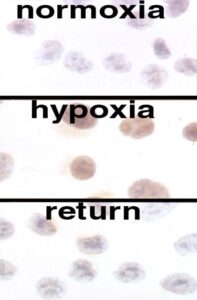
Immunoperoxidase labelling of HIF-1a showing low levels in normoxic tissue culture cells (top), higher levels in hypoxic tissue culture cells (middle) and the rapid loss of this protein on reoxygenation (bottom).
Using the antibodies we generated that recognised HIF-1alpha (and subsequently HIF-2alpha) we were able to show rapid changes in protein abundance as the oxygenation status of cells was altered. We, and others, were then able to show that these changes in protein abundance were mediated through proteasomal degradation of the alpha chains when oxygen was present. Proteins are commonly targeted for proteasomal destruction after they have been tagged with ubiquitin by a specific ubiquitin E3 ligase, so the next step was to try to identify the cognate ubiquitin ligase for HIF-alpha proteins. A variety of pieces of circumstantial evidence led us to the von Hippel Lindau protein (pVHL) as a prime candidate – it had recently been shown to be part of a protein complex with structural similarities to other ubiquitin E3 ligases and patterns of gene expression seen in renal cancer cells lacking pVHL overlapped with those regulated by HIF. Patrick Maxwell spearheaded the work that confirmed that there was an interaction between pVHL and HIF-alpha proteins [The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. PH Maxwell, MS Wiesener, G-W Chang, SC Clifford, EC Vaux, ME Cockman, CC Wykoff, CW Pugh, ER Maher, PJ Ratcliffe. Nature 399 (1999) 271-275. PMID: 10353251]. Emma Vaux, who had come to the lab after working as a renal registrar in Oxford, contributed to this work whilst also continuing the mutagenesis and complementation project outlined earlier that Morwenna Wood had done so much to set up. This ultimately paid dividends, but only after the oxygen-sensing pathway had been elucidated by other means [Selection and analysis of a mutant cell line defective in the hypoxia inducible factor-1 alpha subunit (HIF-alpha): characterisation of HIF-1α dependent and independent gene expression. SM Wood, MS Wiesener, KM Yeates, N Okada, CW Pugh, PH Maxwell, PJ Ratcliffe. Journal of Biological Chemistry 273 (1998) 8360-8368. PMID: 9525945; Selection of mutant CHO cells with constitutive activation of the HIF system and inactivation of the von Hippel-Lindau tumor suppressor. EC Vaux, SM Wood, ME Cockman, LG Nicholls, KM Yeates, CW Pugh, PH Maxwell, PJ Ratcliffe. Journal of Biological Chemistry 276 (2001) 44323-44330. PMID: 11555645].

Emma Vaux after her DPhil viva 2001.
Subsequent work, facilitated by the purchase of a newly developed hypoxia workstation from Andrew Skinn using an MRC equipment grant I obtained, allowed more detailed analysis. Importantly, the workstation allowed us to show that when oxygen was excluded from the protein preparation the interaction was truly oxygen-dependent, that the interaction did lead to ubiquitylation of the HIF-alpha chain and ultimately the precise oxygen dependence of the reactions leading to that interaction. [Hypoxia inducible factor-alpha binding and ubiquitylation by the von Hippel-Lindau tumor suppressor protein. ME Cockman, N Masson, DR Mole, P Jaakkola, G-W Chang, SC Clifford, ER Maher, CW Pugh, PJ Ratcliffe, PH Maxwell. Journal of Biological Chemistry 275 (2000) 25733-25741. PMID: 10823831].
IMAGE FOLLOWS SOON
Co-immunoprecipitation of HIF alpha proteins with VHL protein
The immunoblot on the left shows that both HIF-1 alpha and HIF-2 alpha co-immunoprecipitate with the VHL protein when proteasomal inhibitors are present. Erroneously this blot suggests that the interaction occurs both in the presence and absence of hypoxia. The blot on the left, performed with extracts made using the hypoxia workstation, demonstrates that the apparent interaction in hypoxia is an artefact of re-oxygenation occurring during sample preparation.
Stabilisation of HIF is a key feature of clear cell renal cancers in which the VHL protein is mutated. This determines many of the phenotypic features of these cancer cells but exactly how it contributes to oncogenesis remains unclear.
The next puzzle was to identify what this interaction depended on. David Mole had now joined the lab and worked alongside a visiting Finnish surgeon, Panu Jaakkola, in a large team effort that established that the interaction depended on hydroxylation of one of the prolyl residues I had mutated back in 1996. Initial experiments showed that synthetic peptides containing the prolyl residue did not interact with the VHL protein unless they had been treated with a cell extract in the presence of oxygen, but not in its absence. In a meticulous set of experiments David and Panu were able to show that the essential component(s) of the cell extract were a heat denaturable macromolecule and that the reaction was temperature dependent suggesting it was enzymatic. By chance, Patrick bumped into Chris Schofield whilst they were watching their respective sons playing football. They fell into conversation and the synergies between our respective scientific interests were now much more obvious and an extremely productive collaboration and friendships ensued.
Subsequent work showed the reaction did indeed require the presence of adequate amounts of iron and oxoglutarate in the cell extract. Parallel mass spectrometry showed that in the presence of oxygen the prolyl residue was gaining 16 daltons in weight, compatible with hydroxylation. Critically a synthetic peptide designed with a hydroxyprolyl residue at the correct position would interact with the VHL protein without requiring any pre-treatments – a result reported to me and Peter by Ya Min Tian in memorable phone calls on a Sunday morning! This work was running neck and neck with parallel work in Bill Kaelin’s lab in Boston. Patrick had a rather tense phone call with him, with me listening in, in which it became clear that both groups had arrived at the same answer at the same time and our papers were published back to back in Science as articles. [Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. P Jaakkola, DR Mole, Y-M Tian, MI Wilson, J Gielbert, SJ Gaskell, A von Kriegsheim, HF Hebestreit, M Mukherji, CJ Schofield, PH Maxwell, CW Pugh, PJ Ratcliffe. Science 292 (2001) 468-472. PMID: 11292861; HIFα Targeted for VHL-Mediated Destruction by Proline Hydroxylation: Implications for O2 Sensing. M Ivan, K Kondo, H Yang, W Kim, J Valiando, M Ohh, A Salic, JM Asara, WS Lane and WG Kaelin Jr, Science 292 (2001) 464-8. DOI: 10.1126/science.10598].

David Mole (centre) with Mat Coleman and Matt Cockman enjoying ten pin bowling.

Chris Schofield
Whilst this work had been going on John O’Rourke, and Andrew Epstein, a clinical training fellow had been continuing our interest in oxygen-sensing systems in model organisms by looking for, and finding a HIF system in the nematode worm C. elegans.
IMAGE FOLLOWS SOON
Evolutionary conservation of the HIF-VHL system in C. elegans.
Top panel – a picture of C. elegans nematode worms
Lower panel (left) – immunoblot of C. elegans HIF from wild type and VHL mutant worms living in normoxic (N) and hypoxic (H) conditions. In wild type worms the HIF is hypoxically inducible whereas in VHL mutant worms it is abundant in the presence of oxygen.
Lower panel (right) – RNase protection assay showing that in C. elegans lactate dehydrogenase mRNA shows an induction pattern compatible with it being a HIF-dependent target gene, as it is in mammalian cells.
We were of course very keen to identify the enzyme(s) responsible for oxygen-dependent prolyl hydroxylation of HIF. The timing was perfect since the C. elegans whole genome had just been published. Chris Schofield used his structural knowledge to predict gene sequences in C. elegans that might encode oxoglutarate-dependent dioxygenases. The worm community kept a central depository of mutant worms with defects in each individual gene, Wormbase, curated by Bob Barsted in the USA. So Andrew simply requested worms bearing mutations in the various genes Chris Schofield had listed and tested whether any of them showed defects in HIF regulation. Whilst HIF regulation was preserved in worms bearing mutations in the collagen prolyl hydroxylases worms bearing three different mutant alleles of a gene called Egl-9 all showed defective HIF regulation. Intriguingly, the previously identified phenotype of these mutant worms was a defect in egg laying and tolerance to a toxin produced by Pseudomonas aeruginosa, rather than anything more obviously related to oxygen sensing. Having the genetic sequence for Egl-9 meant it was relatively straightforward to identify the three mammalian homologues, which Jonathan Gleadle playfully labelled as PHD (for prolyl hydroxylase domain) 1-3, on the basis that studies on each enzyme would provide at least one PhD thesis! [C. elegans EGL-9 and mammalian homologs define a family of dioxygenases that regulate HIF by prolyl hydroxylation. ACR Epstein, JM Gleadle, LA McNeill, KS Hewitson, JF O’Rourke, DR Mole, M Mukherji, E Metzen, MI Wilson, A Dhanda, Y-M Tian, N Masson, DL Hamilton, P Jaakkola, R Barstead, J Hodgkin, PH Maxwell, CW Pugh, CJ Schofield, PJ Ratcliffe. Cell 107 (2001) 43-54. PMID: 11595184]
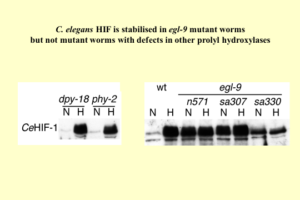
HIF regulation was preserved in worms bearing mutations in the collagen prolyl hydroxylases (dpy-18; phy-2) whereas worms bearing three different mutant alleles (n571; sa307; sa330) of a gene called Egl-9 all showed defective HIF regulation.

Andy Epstein – post-viva celebration
This left open the question of how the activity of the carboxy-terminal transactivation domain of HIF-alpha chains was regulated by hypoxia. Parallel work in Australia and Oxford demonstrated that it was the target of another oxoglutarate-dependent dioxygenase called Factor-inhibiting HIF (FIH). [Hypoxia Inducible Factor (HIF) asparagine hydroxylase is identical to Factor Inhibiting HIF (FIH) and is related to the cupin structural family. KS Hewitson, LA McNeill, MV Riordan, Y-M Tian, AN Bullock, RW Welford, JM Elkins, NJ Oldham, S Bhattacharya, JM Gleadle, PJ Ratcliffe, CW Pugh, CJ Schofield. Journal of Biological Chemistry 277 (2002) 26351-26355. PMID: 12042299; FIH-1 is an asparaginyl hydroxylase enzyme that regulates the transcriptional activity of hypoxia-inducible factor. D Lando , D J Peet, J J Gorman, D A Whelan, M L Whitelaw, R K Bruick. Genes Dev 16(2002) 1466-71.]
The beauty of finding that enzymes were central to hypoxic gene regulation is that they make excellent drug targets, opening opportunities for therapeutic manipulation of this pathway. Subsequent work in pharmaceutical companies has now led to the licensing of HIF hydroxylase inhibitors for the treatment of renal anaemia (Daprodustat, Roxadustat, Vadadustat), and in parallel work a specific inhibitor of HIF-2alpha (Belzutifan) has been licensed for the treatment of renal cancer.
In 2019 the Nobel Committee recognised the combined work of Gregg Semenza, Peter Ratcliffe and Bill Kaelin in understanding cellular oxygen sensing by the award of the Nobel Prize in Medicine or Physiology.

Nobel Prize 2019 – in the lab on the day the news broke. Peter Ratcliffe has his back to the camera whilst in discussion with the team from the BBC.
In the foreground (clockwise from bottom left): Jess McKindrick, Tom Keeley, David Mole, Tammie Bishop, Olivia Lombardi, Sylvie Halim, Gloria Cheng, Chris Pugh, Ya Min Tian, Joanna Lima

Nobel Prize giving 2019 l-r Chris Pugh, Patrick Maxwell, Peter Ratcliffe and David Mole

Nobel Prize giving 2019 l-r Bill Kaelin, Peter Ratcliffe and Gregg Semenza


Nobel Prize giving 2019 – Peter Ratcliffe receiving his award from the King of Sweden
Where are the clinician scientists now?
Nephrologists
Sir Peter Ratcliffe FRS FMedSci
Professor of Clinical Medicine, University of Oxford and Director of Clinical Research, The Francis Crick Institute, London.
Tan Chorh Chuan
Permanent Secretary (National Research and Development), Permanent Secretary (Public Sector Science and Technology Policy and Plans Office), Prime Minister’s Office and Chairman of the Agency for Science, Technology and Research (A*STAR) in Singapore. In addition, he is the Chairman of the Ministry of Health Office for Healthcare Transformation (MOHT) and the Chair of MOH’s Healthier Singapore Implementation Office.
Chris Pugh FMedSci
Professor of Renal Medicine, University of Oxford; Director of Oxford University Clinical Academic Graduate School.
Patrick Maxwell FMedSci
Regius Professor of Physic and Head of the School of Clinical Medicine at the University of Cambridge.
John Firth
Consultant Physician and Nephrologist, Deputy Medical Director and Responsible Officer, Cambridge University Hospitals.
Kai Uwe Eckardt
Professor of Medicine and Head of the Department of Nephrology and Medical Intensive Care at the Charité in Berlin, one of Europe´s largest university hospitals.
Jonathan Gleadle
Consultant Nephrologist at Flinders Medical Centre and Professor of Medicine at Flinders University.
Neil Bacon
Chief Strategy and Transformation Officer for NHS Dorset Integrated Care Board.
Morwenna Wood
Director of Medical Education, NHS Fife; Honorary Professor, University of St Andrews.
Emma Vaux
Consultant Nephrologist and General Physician at the Royal Berkshire NHS Foundation Trust and global vice president at the Royal College of Physicians.
David Mole
Professor of Renal Medicine, University of Oxford.
Other specialties
Ben Ebert
President and CEO of Dana-Farber Cancer Institute and the George P. Canellos, MD and Jean S. Canellos Professor of Medicine at Harvard Medical School.
Andrew Epstein
Consultant Cardiologist in Perth, W. Australia.
Last Updated on December 12, 2025 by John Feehally