Polycystic Kidney Disease
ADPKD is a common, dominant monogenic disorder that causes 5-10% of kidney failure worldwide. Thirty years ago, the most common gene causing autosomal dominant polycystic kidney disease (ADPKD), PKD1, was identified. Scientists in the UK made major contributions to the early stages of unraveling the cause of this complex disorder.
Contents
Localisation of the PKD1 gene
In the dark ages before the Human Genome Projects gave us the sequence of the whole genome and even before a good genetic map or DNA clones covering the genome were available, the first step to identifying a monogenic disease gene was to find roughly in the genome where the gene lies. The central protagonist of this first stage of the story was Steve Reeders, a precocious clinical fellow working with Sir David Weatherall and Doug Higgs, in the Nuffield Department of Medicine at the John Radcliffe Hospital in Oxford.
The strategy was to collect DNA samples from patients with ADPKD from large families and then look for polymorphic markers from across the genome that segregate with disease in these families. As luck would have it, Higgs had recently found a highly polymorphic marker next to the a-globin gene, located on the short arm (p) of chromosome 16. This marker, 3’HVR, out of the hundreds of markers throughout the genome that could have been tested, was found to be closely linked (likely within 5Mb) to the ADPKD gene in the collected families. This important first step to identifying this gene was published in Nature in 1985.
In the next few years, groups worldwide including the Oxford group, identified and tested further markers and localised the gene close to the tip of chromosome 16p. During that time, a postdoctoral fellow working at Harvard, Peter Harris, started to collaborate with the Reeders’ group to localise the gene, and at the end of 1987 he moved to Oxford to work on this project with Reeders. However, soon after he arrived, Reeders moved to Yale, but Harris continued to work on the project, now in the Institute of Molecular Medicine (IMM, later named the Weatherall IMM), Oxford.
Continued mapping efforts indicated that the disease was genetically heterogeneous. There was a group of ADPKD families with a gene defect not linked to 16p, these were named PKD2, and the 16p gene was named PKD1. By 1992 these mapping efforts had narrowed PKD1 to a 600kb region. Also, during this period, Peter Harris started a collaboration (facilitated by Sir Peter Ratcliffe (Nephrologist, Nuffield Professor of Medicine at Oxford, and Nobel laureate) with Chris Winearls, the head of the Renal Unit, Churchill Hospital, Oxford, to collect DNA samples and establish cell lines from tens and then hundreds of ADPKD patients, work enabled by a study coordinator, Carol Strong. Although the region containing the gene was quite small (by whole genome standards), identification of the specific gene remained elusive, partly because many genes are localized to this GC-nucleotide rich region of the genome.
It’s near TSC2
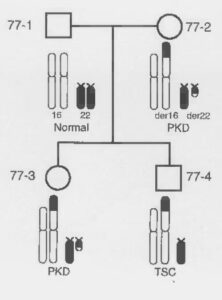
The next lucky break in this story came with a call to Peter Harris from Julian Sampson (Department of Medical Genetics, Cardiff.) Sampson was interested in another dominant disease, tuberous sclerosis (TSC), that also had a locus recently mapped to the tip of 16p (TSC2). In addition, he had heard of a Portuguese family being studied by Isabel Cotdeiro ( Genetics Unit, Hospital Santa Maria, Lisbon) that had ADPKD and TSC in different family members. Even more remarkably, this family segregated with a chromosomal translocation including the 16p region of interest and chromosome 22. Further analysis of this family confirmed that the mother and daughter with the balanced translocation had ADPKD while the son with an unbalanced karyotype, missing the tip of 16p and part of chromosome 22, had TSC.
A possible explanation was that TSC2 gene was localized in the deleted region of 16p, while the breakpoint of the translocation disrupted the PKD1 gene itself. Although this is rarely how things work out in science, in this case the hypothesis turned out to be true! Harris and Sampson then involved their collaborators on ADPKD, Martijn Breuning, and TSC, Dicky Halley, from The Netherlands to form a European Consortium to identify both genes.
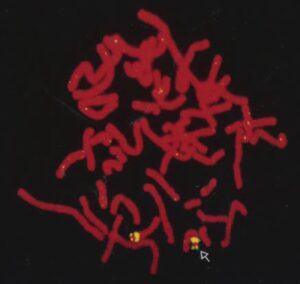
A frantic period followed – refining the breakpoint of the translocation using existing and newly identified clones by postdoc Chris Ward and PhD student Jim Hughes. The methods employed were fluorescence in situ hybridization (FISH) with an Oxford collaborator, Veronica Buckle; somatic cell hybrids with other translation breakpoints in this region; and field inverted gel electrophoresis (FIGE). The breakpoint was first localized to a 250kb region, refined by FIGE to 60kb, and eventually the precise breakpoint was identified. At the same time, screening of cDNA libraries for transcripts possibly encoded by this region identified several candidate genes. One of these, just distal to the translocation breakpoint was found from the identification of point mutations in TSC patients to be TSC2. A finding published in Cell by the European Consortium in 1993 with Julian. Sampson as the corresponding author.
Sequencing PKD1: huge and complex
Identifying and obtaining the full sequence of the PKD1 gene proved a little more difficult. A gene that was shown to straddle the translocation breakpoint was identified but appeared from Northern blot analysis to encode a very large transcript (we now know it is more than 16kb in size). Furthermore, the position of the breakpoint lay in a segmentally duplicated region of the genome with multiple (we now know there are six) copies of a highly homologous region located more proximally on 16p. This duplicate region also contained transcribed pseudogenes with a high level of sequence similarity to PKD1. For these reasons, the initial transcript characterization and analysis for disease causing pathogenic variants in ADPKD patients focused on the 3’ part of this gene lying between TSC2 and the start of the duplicated region (we now know to be a coding region of 2.6kb). These studies by Belen Peral, Vicki Gamble and Sandra Thomas identified two multiexon deletions by FIGE and Northern blotting, and a canonical spicing mutation by analysis of the transcript.
At that point the decision was made to try to publish this discovery even though the whole gene transcript had not been identified and the number of disease-causing variants was limited. This gamble paid off with the initial identification of PKD1 published by the European Consortium in Cell in June 1994 with Peter Harris as the corresponding author. Harris PKD 1 gene Cell 1994. The finding that two major dominant disease genes, TSC2 and PKD1, lay with their 3’ ends just 60bp apart was another amazing part of this story. This juxtaposition also explained occasional TSC patients who had severe, early onset PKD (TSC patients sometimes have a few kidney cysts). The explanation was contiguous deletions disrupting PKD1 and TSC2, a finding published in Nature Genetics in 1994.
Completing the full sequence
Although the PKD1 gene was now identified, the work was not over because the full transcript needed to be cloned and sequenced. The Oxford group employed a somatic cell hybrid that just contained PKD1 but not the pseudogene encoding region of chromosome 16 to define the bona fide PKD1 transcript. The result of this was the identification of a 12906bp encoding region for a 4302 amino acid protein, named polycystin 1 (PC1), published in 1995 in Nature Genetics by the Harris group (similar finding were published by an International Consortium, including Reeders in the same year). The achievement was described in MRC News (pdf, 1.2Mb).
Understanding the structure and function of PKD1 and PKD2
PC1 is a unique protein with 11 transmembrane domains, a large extracellular region encoding 16 copies of a domain now called the PKD Repeat, and several other recognized protein and carbohydrate interaction domains. The large size of the protein has made understanding its function difficult, but Chris Ward and Albert Ong (who came to Oxford as a Senior Fellow just after the gene was identified and is now Professor of Nephrology at the University of Sheffield) have made major contributions in this area. In 1996, the group of Stefan Somlo (Albert Einstein, New York) identified the second major ADPKD gene, PKD2. Its identification was facilitated by homology between the PC2 and PC1 proteins.
Since the identification of these genes important discoveries have been made that help understand the disease. Over 2000 different pathogenic variants have been described, with no common mutation; PC1 and PC2 form a complex that likely has channel function; and the primary cilium, singularly found on most cell types in the body, including epithelial cells where it protrudes from the apical surface, is a likely functional site of the PC complex. Multiple pathways have been implicated in ADPKD pathogenesis, including dysregulated cAMP signalling, that has led to the regulatory approval of the first drug specifically for ADPKD. However, 30 years since PKD1 was identified, many questions remain, including the primary function of the PC-complex.
This story, with more than one fortuitous turn, continues. Complete understanding of ADPKD disease pathogenesis, and treatments that fully halt disease progression are still to be found. However, the story illustrates several points that are emblematic of scientific discovery: serendipity often plays a role, but keen observation and hard work are needed to turn these opportunities into scientific breakthroughs.
Authorship
Peter Harris. First posted Jan 2024.
Last Updated on September 12, 2024 by neilturn